




Uncover the science behind it: pmc.ncbi.nlm.nih.gov/articles/PMC...

Uncover the science behind it: pmc.ncbi.nlm.nih.gov/articles/PMC...
Google AI, in collaboration with the UC Santa Cruz Genomics Institute, has introduced DeepPolisher, a cutting-edge deep learning tool designed to…
Google AI, in collaboration with the UC Santa Cruz Genomics Institute, has introduced DeepPolisher, a cutting-edge deep learning tool designed to…


@kseniakrasileva.bsky.social will be there presenting on some of our latest work that was submitted to BioRxiv today. Teaser: Plant immunity + large language models will transform receptor discovery and engineering for disease resistance. 🌱🚀
@kseniakrasileva.bsky.social will be there presenting on some of our latest work that was submitted to BioRxiv today. Teaser: Plant immunity + large language models will transform receptor discovery and engineering for disease resistance. 🌱🚀
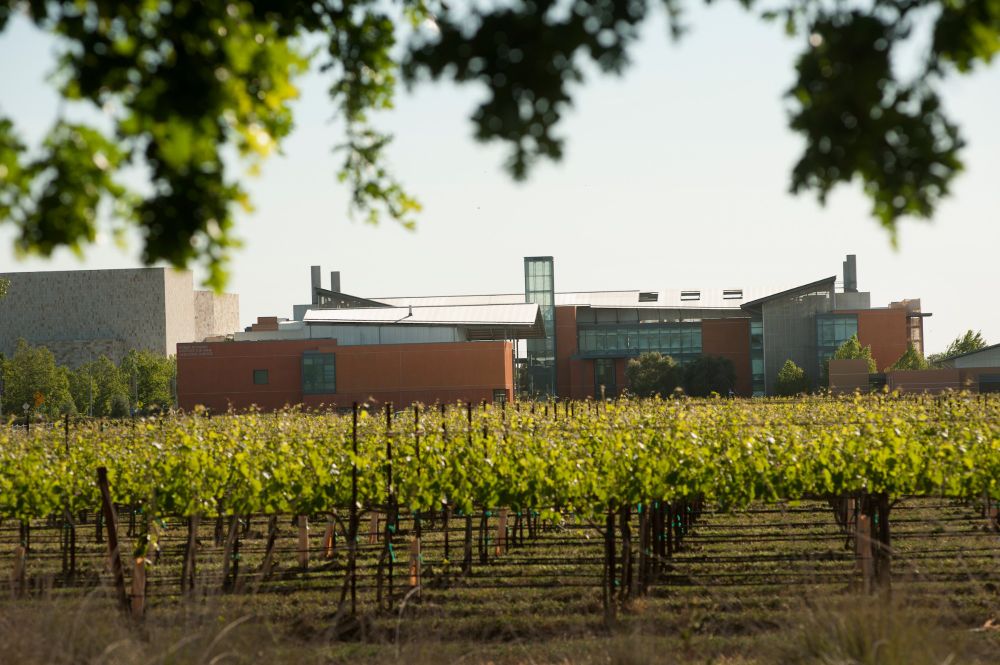


What are we doing?
What are we doing?


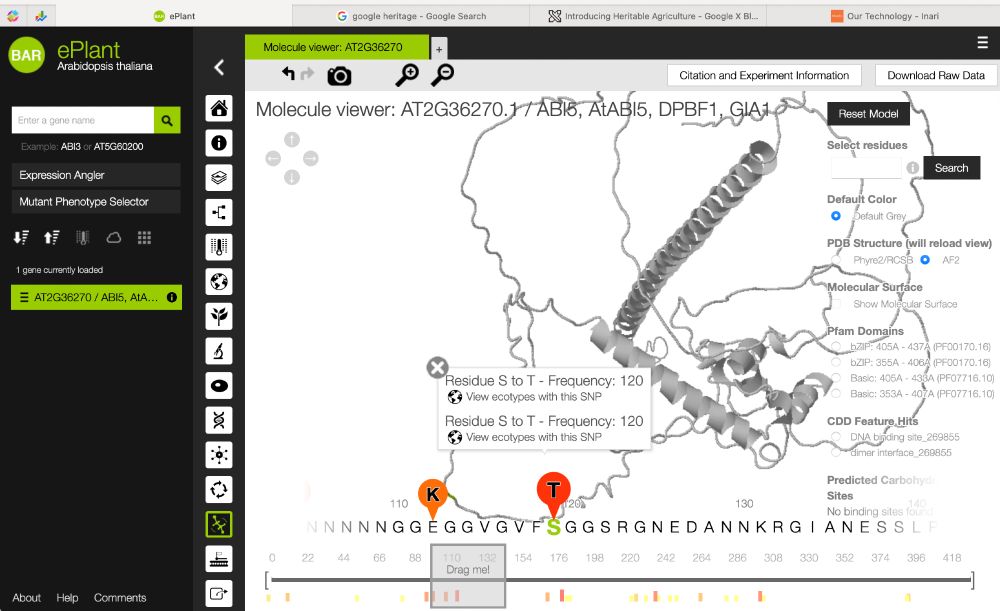
#PlantScience @bar-plantbio.bsky.social
If so, you might this little project: github.com/rrwick/conda...
If so, you might this little project: github.com/rrwick/conda...
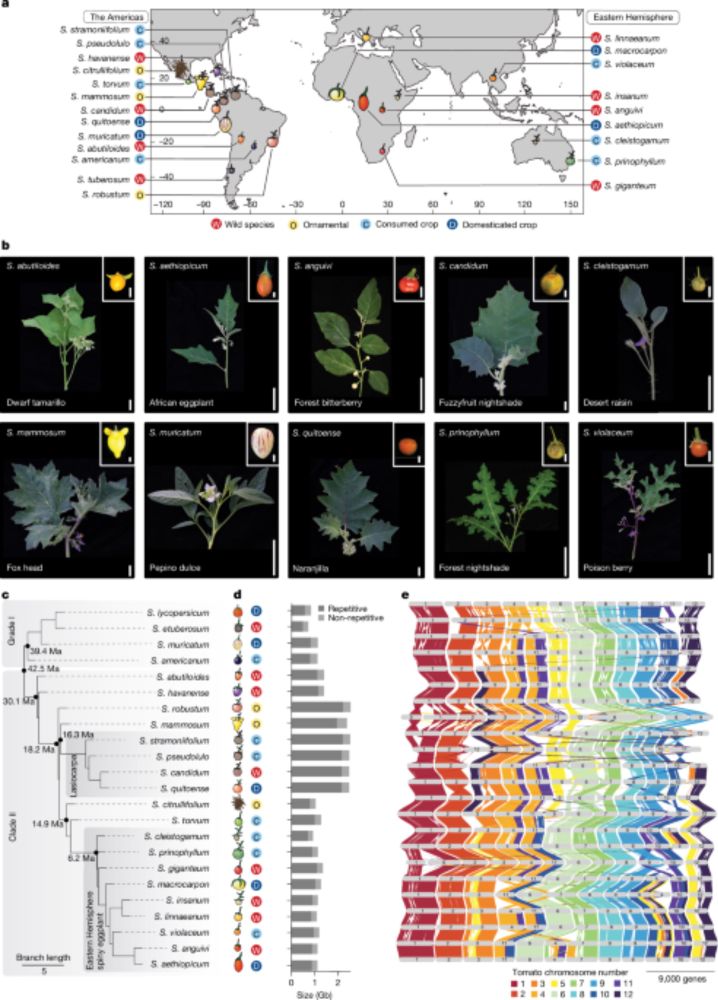
www.nature.com/articles/s41...

www.nature.com/articles/s41...





▶️ www.science.org/doi/10.1126/...
#Genetics

▶️ www.science.org/doi/10.1126/...
#Genetics



