ICYMI, also happy to discuss our new Aardvark tool for benchmarking :)
Aardvark: Sifting through differences in a mound of variants
GitHub: github.com/PacificBiosc...
Some highlights in this thread:
1/N

doi.org/10.1093/bioi...

doi.org/10.1093/bioi...
The paper describes STRchive as a resource to improve the diagnosis of tandem repeat disorders, then goes beyond it to consider what can be learned about childhood onset and population prevalence of these diseases.
🖥️ 🧬
link.springer.com/article/10.1...
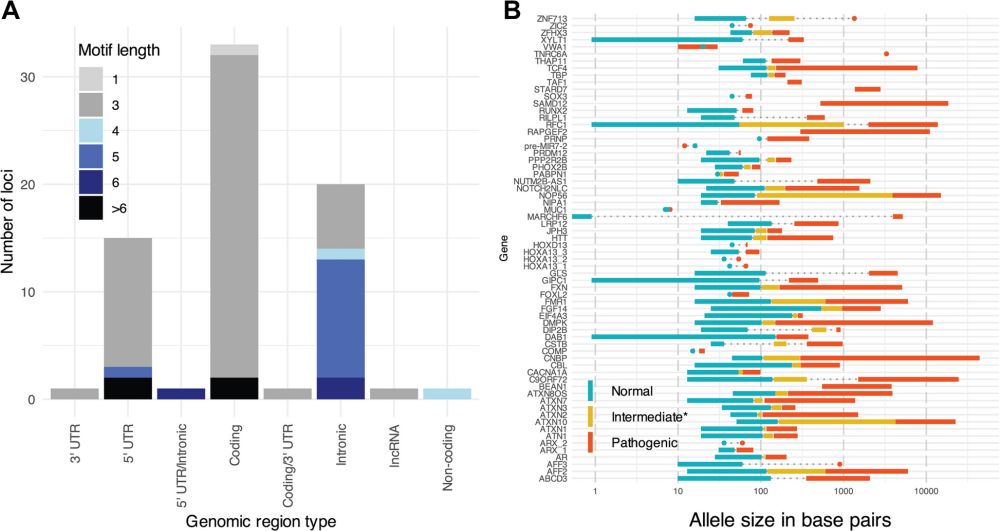
The paper describes STRchive as a resource to improve the diagnosis of tandem repeat disorders, then goes beyond it to consider what can be learned about childhood onset and population prevalence of these diseases.
🖥️ 🧬
link.springer.com/article/10.1...
Check out the database resource, as STRchive "streamlines TR variant interpretation at disease-associated loci."
strchive.org
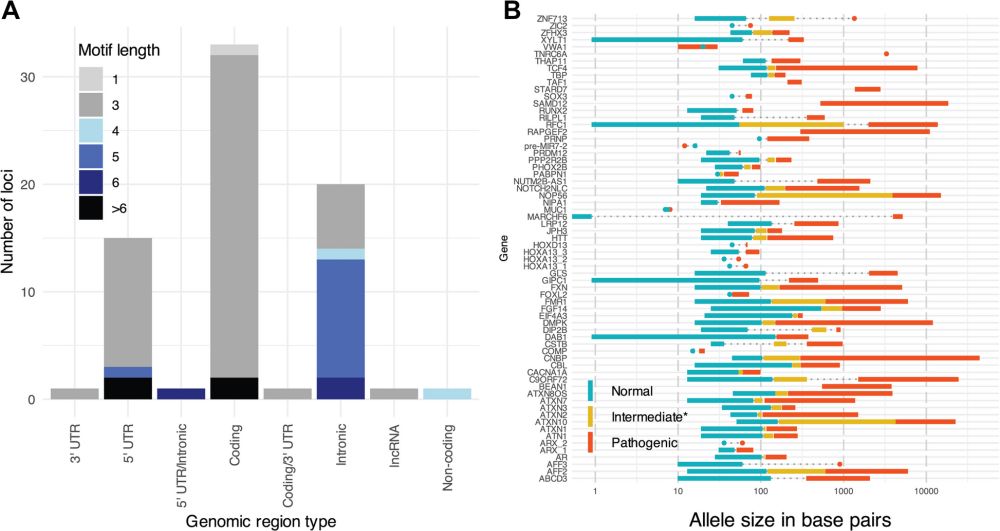
Check out the database resource, as STRchive "streamlines TR variant interpretation at disease-associated loci."
strchive.org
Learn what surprising mechanism was uncovered, the potential impact on therapeutic developments, & how #PacBio played a role. www.cell.com/cell/fulltex...

Learn what surprising mechanism was uncovered, the potential impact on therapeutic developments, & how #PacBio played a role. www.cell.com/cell/fulltex...

www.cell.com/ajhg/abstrac...

www.cell.com/ajhg/abstrac...
jobs.ac.uk/job/DLG312/bio…
jobs.ac.uk/job/DLG312/bio…
Link for application:
candidate.hr-manager.net/ApplicationI...

Link for application:
candidate.hr-manager.net/ApplicationI...
A resource for tandem repeats associated with Mendelian disease. We have resigned the website, added new loci, streamlined our data for easier reuse, added more detailed citations, presented population frequency data and more!

A resource for tandem repeats associated with Mendelian disease. We have resigned the website, added new loci, streamlined our data for easier reuse, added more detailed citations, presented population frequency data and more!
Pre-print: doi.org/10.1101/2024...
Repo: github.com/PacificBiosc...

Pre-print: doi.org/10.1101/2024...
Repo: github.com/PacificBiosc...
Pre-print: doi.org/10.1101/2024...
Repo: github.com/PacificBiosc...


If you were interested but missed it: www.pacb.com/wp-content/u...
If you were interested but missed it: www.pacb.com/wp-content/u...
Import bluesky as bs
del x
bs.activate()
Import bluesky as bs
del x
bs.activate()
www.pacb.com/wp-content/u...
www.pacb.com/wp-content/u...

