Developing advanced methods for atomistic simulations.

pubs.acs.org/doi/full/10....

pubs.acs.org/doi/full/10....
pubs.acs.org/doi/10.1021/...
pubs.acs.org/doi/10.1021/...
Paper: arxiv.org/abs/2505.16476
Code: github.com/NeuralTSNE

Paper: arxiv.org/abs/2505.16476
Code: github.com/NeuralTSNE

nawa.gov.pl/en/programy-...
nawa.gov.pl/en/programy-...
doi.org/10.1063/5.02...
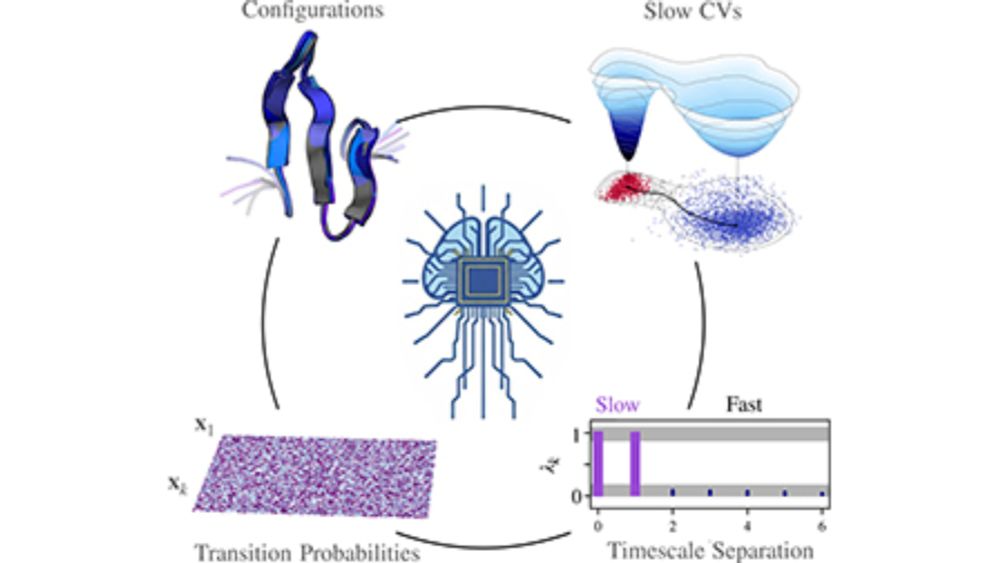
doi.org/10.1063/5.02...
on her 1st first-author paper! arxiv.org/abs/2412.20868
Many thanks to Haochuan Chen, Luke Evans, Luigi Bonati, and Omar Valsson for their great feedback!

on her 1st first-author paper! arxiv.org/abs/2412.20868
Many thanks to Haochuan Chen, Luke Evans, Luigi Bonati, and Omar Valsson for their great feedback!
For anyone interested in spectral maps.
I made a typo in one equation in the original paper published in JCPL. This has now been fixed; see the accompanying correction note.
pubs.acs.org/doi/full/10....
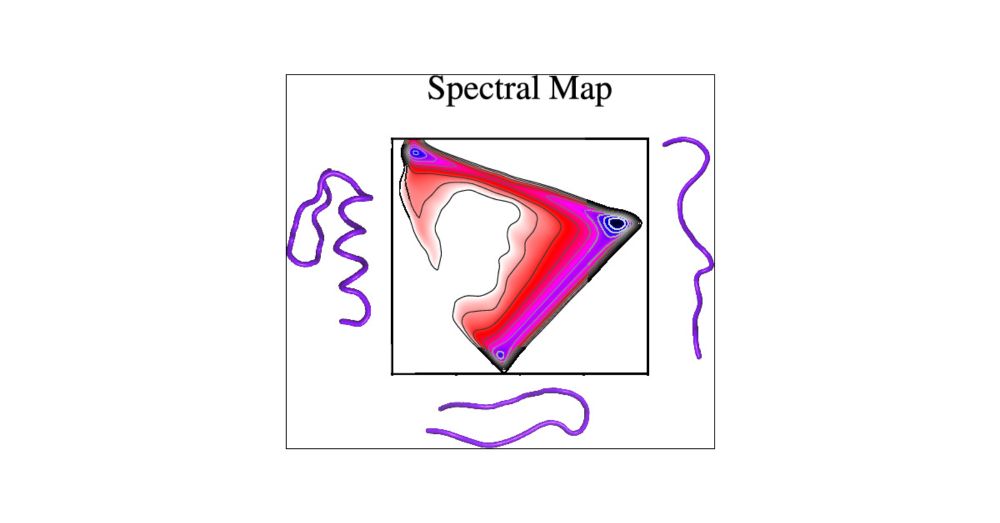
For anyone interested in spectral maps.
I made a typo in one equation in the original paper published in JCPL. This has now been fixed; see the accompanying correction note.
pubs.acs.org/doi/full/10....
Implementation: github.com/jakryd/plume...
More details about PLUMED Tutorials in our collaborative work: arxiv.org/abs/2412.03595
Implementation: github.com/jakryd/plume...
More details about PLUMED Tutorials in our collaborative work: arxiv.org/abs/2412.03595

Mail or DM me for more information.
fizyka.umk.pl/en/prom-eng/

Mail or DM me for more information.
fizyka.umk.pl/en/prom-eng/


