
https://www.genomes-in-time.org
🌱 Full Professor in Stress Resilience of Plants (W3 tenured)
📌What are we looking for? Someone working with us strategically at the interface of plant biology/ physiology/ resilience
#academicjobs #facultyjobs #PlantJobs
1/4

🌱 Full Professor in Stress Resilience of Plants (W3 tenured)
📌What are we looking for? Someone working with us strategically at the interface of plant biology/ physiology/ resilience
#academicjobs #facultyjobs #PlantJobs
1/4
- very low, scattered abundance with barely intact genes in Hydnora
- extreme amplification in sister genus Prosopanche (> 15 % of genome; is this the highest value ever seen?)
- extreme sequence divergence
---> we wonder...
(4/6)
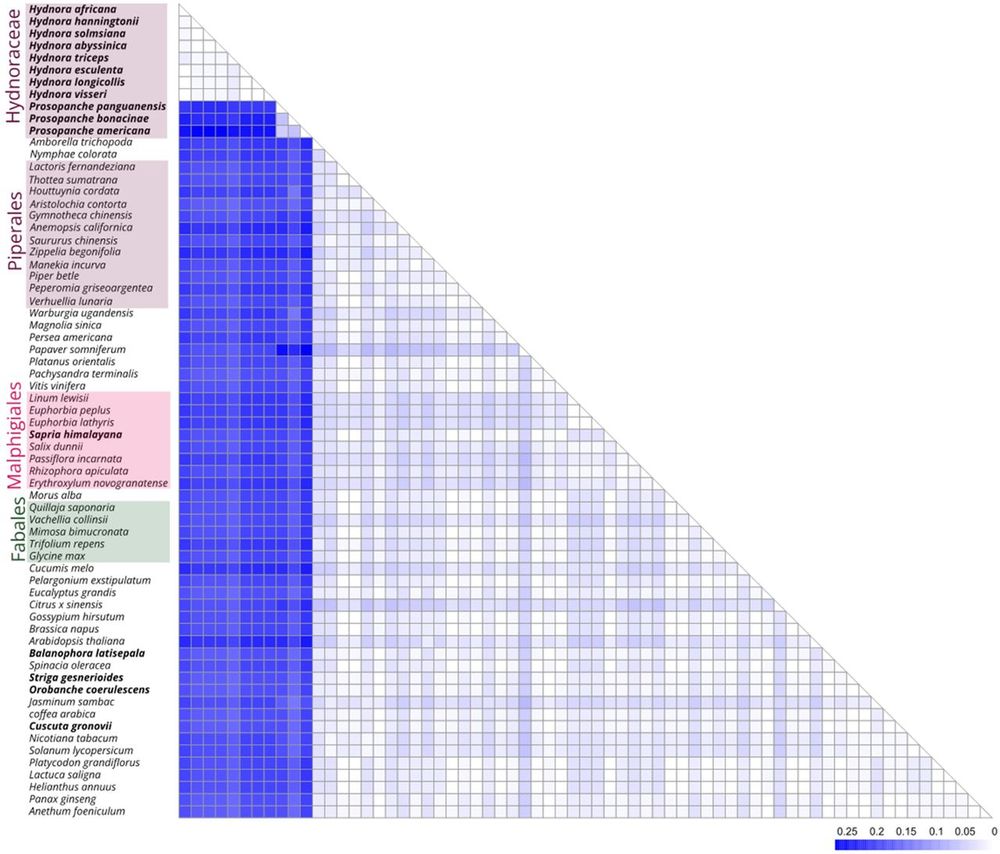
- very low, scattered abundance with barely intact genes in Hydnora
- extreme amplification in sister genus Prosopanche (> 15 % of genome; is this the highest value ever seen?)
- extreme sequence divergence
---> we wonder...
(4/6)
1) Hydnora with 8 sp.:
a lot of gypsy retrotransposons; nearly no 5S rDNA; similar compositions
2) Prosopanche with 3 sp.:
very different compositions; high amplification of an En/Spm DNA transposon (P. bonacinae) and the 5S (P. panguanensis)
(3/6)

1) Hydnora with 8 sp.:
a lot of gypsy retrotransposons; nearly no 5S rDNA; similar compositions
2) Prosopanche with 3 sp.:
very different compositions; high amplification of an En/Spm DNA transposon (P. bonacinae) and the 5S (P. panguanensis)
(3/6)
📚 Read here: www.biorxiv.org/content/10.1...
🌱We look into the repeat landscapes of 11 species from an iconic holoparasitic plant family, the Hydnoraceae.
(1/6)

📚 Read here: www.biorxiv.org/content/10.1...
🌱We look into the repeat landscapes of 11 species from an iconic holoparasitic plant family, the Hydnoraceae.
(1/6)
🌱Visiting the #PlantScience mega-congress #IBC2024 in Madrid?
Let's meet, let's greet, let's connect! And let us know, if you plan to join, so that we can count you in (forms.gle/sDmBA1GScN2Q...).

🌱Visiting the #PlantScience mega-congress #IBC2024 in Madrid?
Let's meet, let's greet, let's connect! And let us know, if you plan to join, so that we can count you in (forms.gle/sDmBA1GScN2Q...).
Attached are some AI images that I had asked in Mucha style ... the AI has mastered it perfectly, i think



Attached are some AI images that I had asked in Mucha style ... the AI has mastered it perfectly, i think
9/14

9/14
All chromosomes have different make-ups and hence are more or less unique. There are also no clearly dominant satellite DNAs. Due to their patchy appearance, we call them "patchwork chromosomes".
8/14

All chromosomes have different make-ups and hence are more or less unique. There are also no clearly dominant satellite DNAs. Due to their patchy appearance, we call them "patchwork chromosomes".
8/14
Only few satellite DNAs are amplified and these are the building blocks of the chromosomes in one of three ways.
7/14

Only few satellite DNAs are amplified and these are the building blocks of the chromosomes in one of three ways.
7/14
In the primary beet gene pool, two satellite DNAs are highly amplified and homogenized. Those two are the building blocks of all beet chromosomes in the primary gene pool.
6/14

In the primary beet gene pool, two satellite DNAs are highly amplified and homogenized. Those two are the building blocks of all beet chromosomes in the primary gene pool.
6/14
5/14

5/14
As so often, larger genome sizes can be explained with more repetitive DNAs (A). Also, more DNA --> more retrotransposons (B). But some fractions (C; D), are not so easily explained. We look more deeply into the satellite DNAs (D)...
4/14

As so often, larger genome sizes can be explained with more repetitive DNAs (A). Also, more DNA --> more retrotransposons (B). But some fractions (C; D), are not so easily explained. We look more deeply into the satellite DNAs (D)...
4/14
- across all three gene pools
- across the two botanical sister genera Beta & Patellifolia
- across all sections and major species, including polyploids
2/14


- across all three gene pools
- across the two botanical sister genera Beta & Patellifolia
- across all sections and major species, including polyploids
2/14
🧬We find that repetitive DNAs evolved differently in the gene pools, marking speciation and crossing boundaries.
doi.org/10.1111/tpj....
1/14

🧬We find that repetitive DNAs evolved differently in the gene pools, marking speciation and crossing boundaries.
doi.org/10.1111/tpj....
1/14
Now, you can read about this work in our preprint:
doi.org/10.1101/2023...

Now, you can read about this work in our preprint:
doi.org/10.1101/2023...
github.com/crimBubble/r...

github.com/crimBubble/r...
🧬🖥️📈find out: doi.org/10.1101/2023...
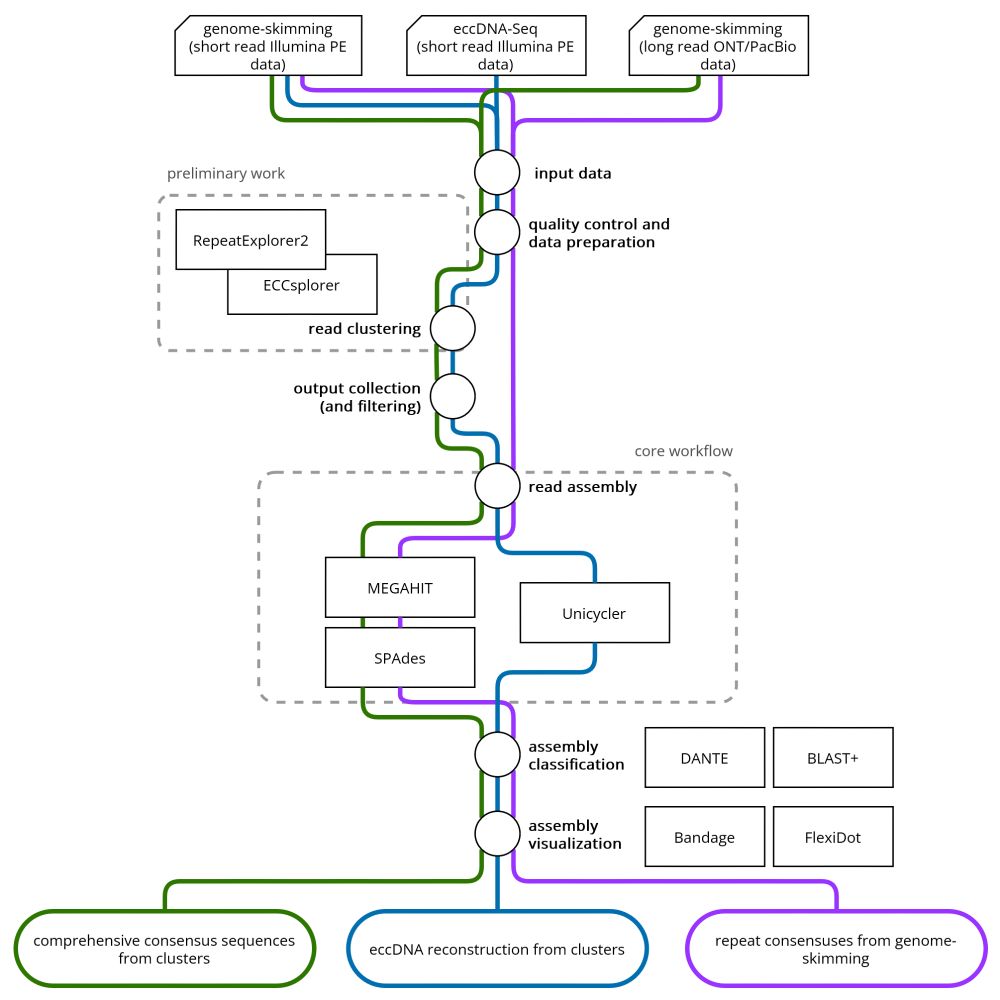
🧬🖥️📈find out: doi.org/10.1101/2023...
I am a biologist & plant genomes and chromosomes are my specialty. I am fascinated by the large, repetitive part of the genome that underlies much of the variability that we can observe.
Just look at the apple 🍏🍎!

I am a biologist & plant genomes and chromosomes are my specialty. I am fascinated by the large, repetitive part of the genome that underlies much of the variability that we can observe.
Just look at the apple 🍏🍎!
Do you enjoy looking at plant chromosomes, but do not know how? Do not worry!
🔎🔬
Sònia Garcia and me worked for two years to bring together exactly 100 authors for a Plant Cytogenetics and Cytogenomics protocol collection!
🧬💚
link.springer.com/book/10.1007...

Do you enjoy looking at plant chromosomes, but do not know how? Do not worry!
🔎🔬
Sònia Garcia and me worked for two years to bring together exactly 100 authors for a Plant Cytogenetics and Cytogenomics protocol collection!
🧬💚
link.springer.com/book/10.1007...

