
(8/)

(8/)
(7/)

(7/)
Modularity adds local structural motifs (i.e., triangles and diamonds), which can increase or decrease the effects of trans-eQTLs (depending on how many genes are activators).
(6/)

Modularity adds local structural motifs (i.e., triangles and diamonds), which can increase or decrease the effects of trans-eQTLs (depending on how many genes are activators).
(6/)
These networks tend to have modular groups, master regulators, a high proportion of activators, and a fixed ratio of sparsity and regulatory strength.
(5/)

These networks tend to have modular groups, master regulators, a high proportion of activators, and a fixed ratio of sparsity and regulatory strength.
(5/)
Here, we use a linear model of expression on randomly generated DAGs, with the goal of finding GRNs that match the observed distribution of cis-h2 fraction.
(4/)

Here, we use a linear model of expression on randomly generated DAGs, with the goal of finding GRNs that match the observed distribution of cis-h2 fraction.
(4/)
Data from pubmed.ncbi.nlm.nih.gov/31558840/
(2/)
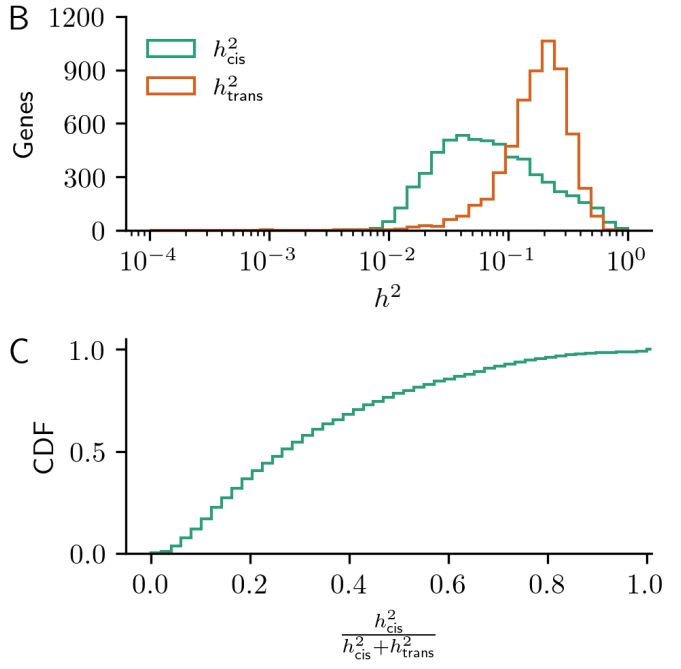
Data from pubmed.ncbi.nlm.nih.gov/31558840/
(2/)

