
@caina89.bsky.social !!! Thank you for this amazing journey!!! 🤩
@caina89.bsky.social !!! Thank you for this amazing journey!!! 🤩
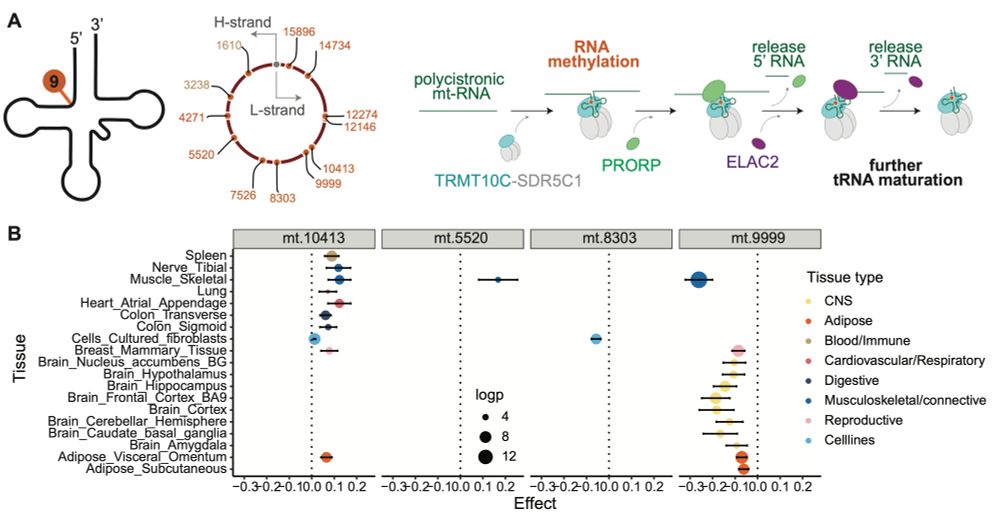
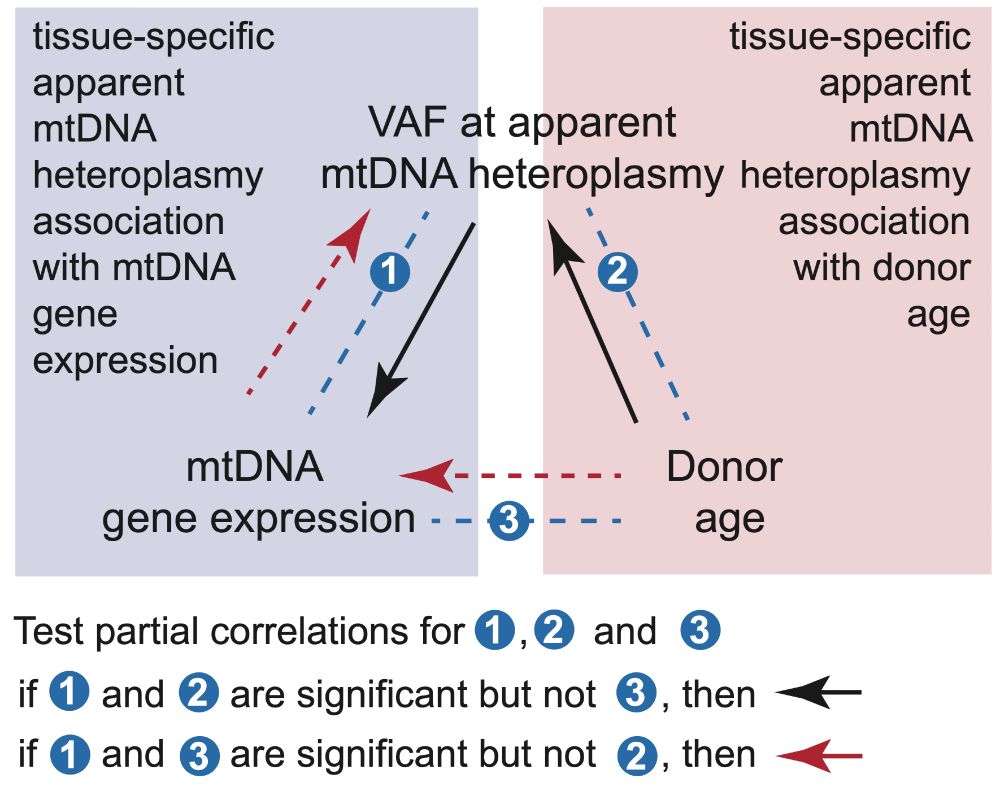

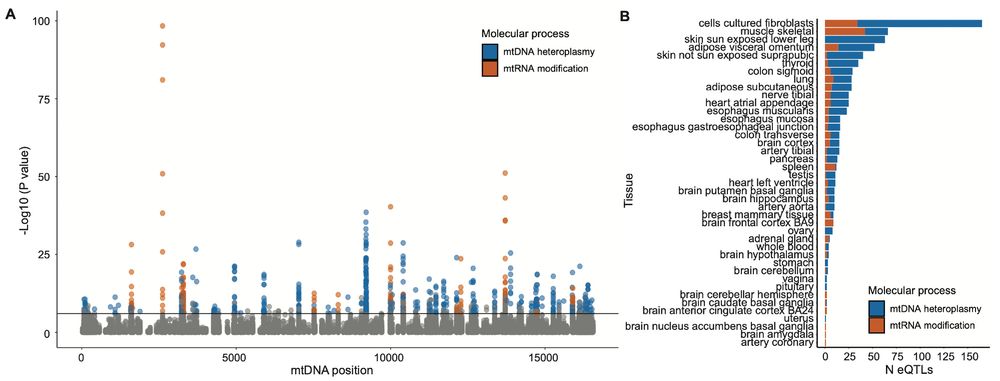
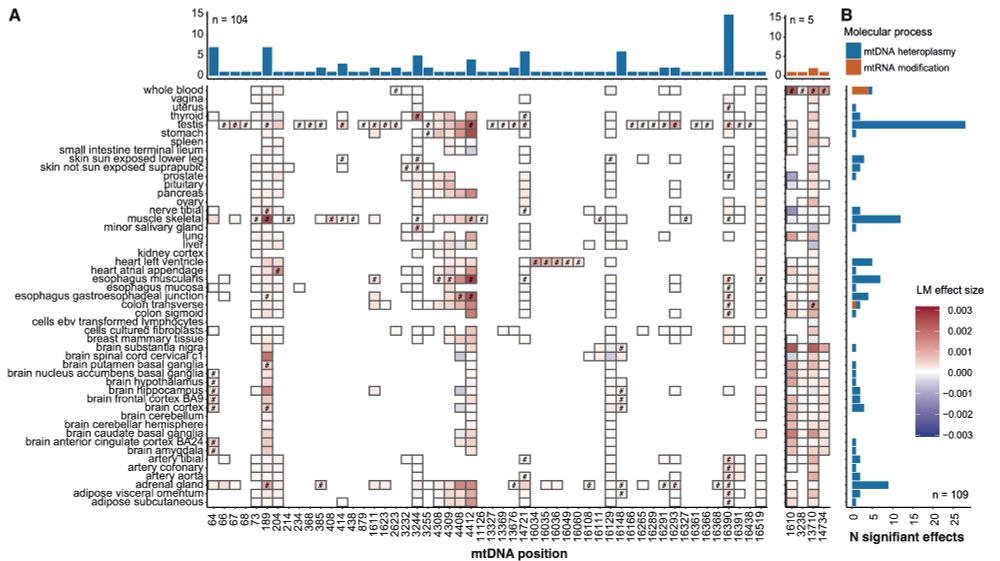
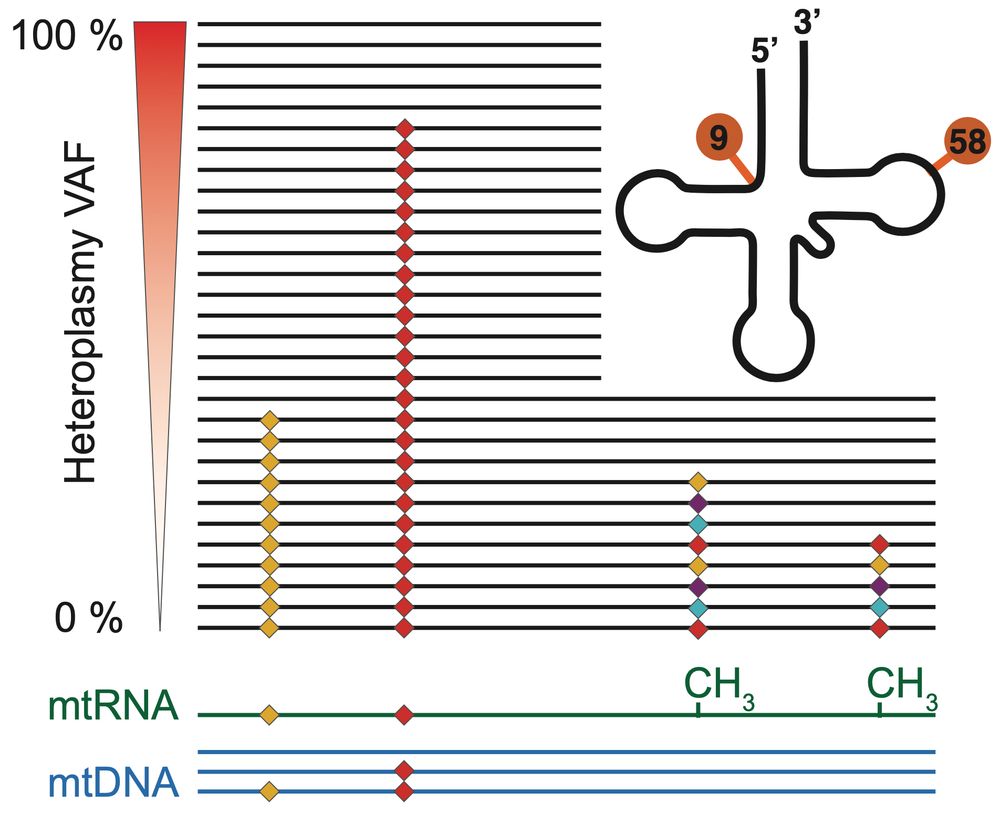

transcribed to RNA and how much are mtRNAs modified post-transcription? We investigate how these variations change with age, and how they affect mtDNA gene expression in a tissue-specific manner.
transcribed to RNA and how much are mtRNAs modified post-transcription? We investigate how these variations change with age, and how they affect mtDNA gene expression in a tissue-specific manner.

