Interests: bioinformatics, cheminformatics, machine learning, Bayesian stats, science fiction and chihuahuas.
Why does this matter? Reproducibility & fair comparison have been lacking—until now.
Paper: arxiv.org/abs/2502.12479 | Repo: github.com/blt2114/Moti...
A thread ⬇️
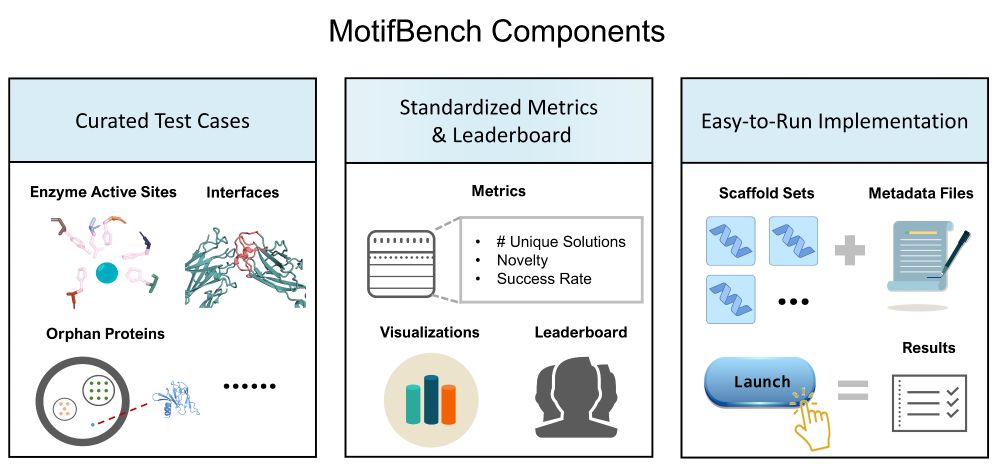
Why does this matter? Reproducibility & fair comparison have been lacking—until now.
Paper: arxiv.org/abs/2502.12479 | Repo: github.com/blt2114/Moti...
A thread ⬇️
if you're interested in the evaluation of conformational accuracy of structure prediction methods, take a look at our first stab at a systematic conformational benchmark in the NP3 technical report below! 🧵
www.iambic.ai/post/np3-tec...
if you're interested in the evaluation of conformational accuracy of structure prediction methods, take a look at our first stab at a systematic conformational benchmark in the NP3 technical report below! 🧵
www.iambic.ai/post/np3-tec...
arxiv.org/abs/2412.02889

arxiv.org/abs/2412.02889
Our answer: They’re two sides of the same coin. We wrote a blog post to show how diffusion models and Gaussian flow matching are equivalent. That’s great: It means you can use them interchangeably.

Our answer: They’re two sides of the same coin. We wrote a blog post to show how diffusion models and Gaussian flow matching are equivalent. That’s great: It means you can use them interchangeably.
1/7
openreview.net/forum?id=H7m...

1/7
openreview.net/forum?id=H7m...
In "Learning on compressed molecular representations" Jan Weinreich and I looked into whether GZIP performed better than Neural Networks in chemical machine learning tasks. Yes, you've read that right.
TL;DR: Yes, GZIP can perform better than baseline GNNs and MLPs. It can ..

december SAGIM event was announced!
looking forward to connecting with fellow comp chemists and modelers over craft beers at new english brewery this december 5th - drop by between 4-7pm
www.meetup.com/southern-cal...

december SAGIM event was announced!
looking forward to connecting with fellow comp chemists and modelers over craft beers at new english brewery this december 5th - drop by between 4-7pm
www.meetup.com/southern-cal...
“if you or someone you know thinks they can actually predict PPIs, prove it. Here is a link to our blinded protein sequence data”
dickinsonlab.uchicago.edu/ppi-challenge
“if you or someone you know thinks they can actually predict PPIs, prove it. Here is a link to our blinded protein sequence data”
dickinsonlab.uchicago.edu/ppi-challenge
arstechnica.com/science/2024...

arstechnica.com/science/2024...


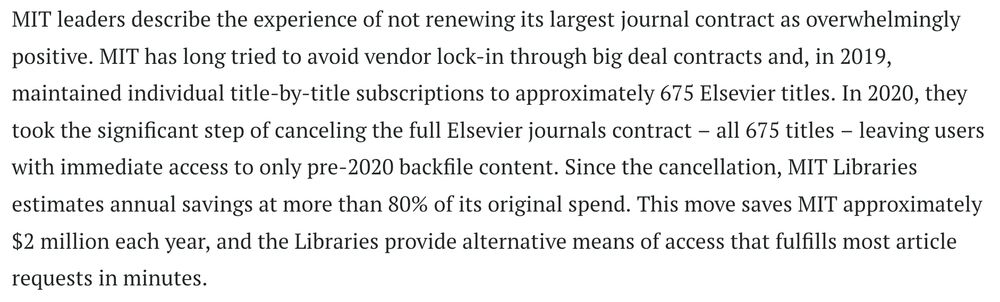
practicalcheminformatics.blogspot.com/2024/05/gene...

practicalcheminformatics.blogspot.com/2024/05/gene...
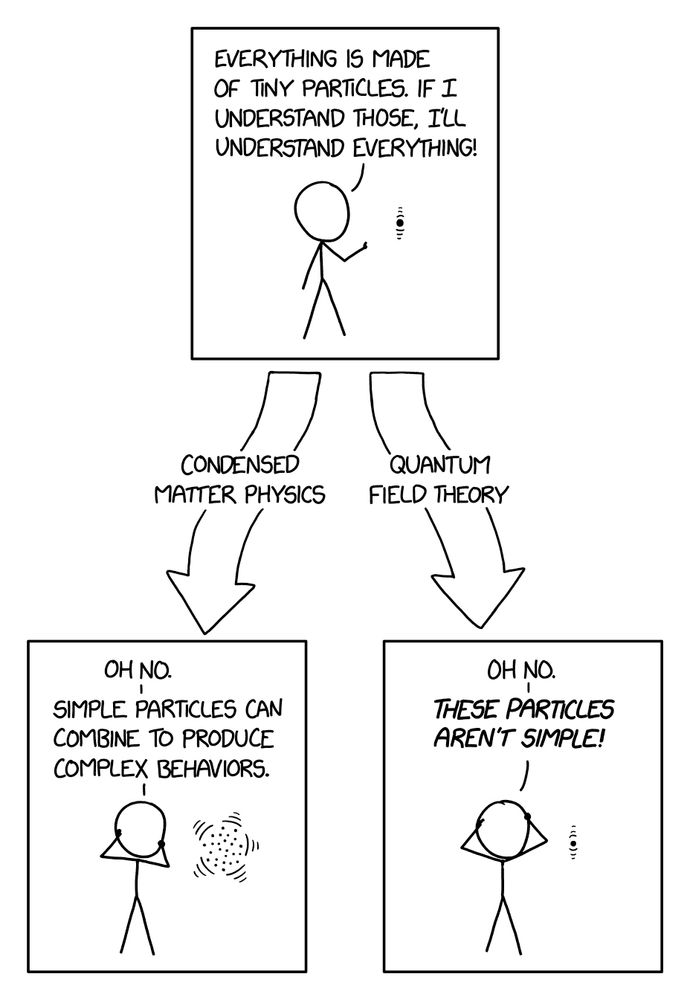
www.biorxiv.org/content/10.1...
www.biorxiv.org/content/10.1...
🧬🧶
📄 www.biorxiv.org/content/10.1...
💾 github.com/steineggerla...
🕸️ search.foldseek.com

🧬🧶
📄 www.biorxiv.org/content/10.1...
💾 github.com/steineggerla...
🕸️ search.foldseek.com

github.com/openmm/spice...

github.com/openmm/spice...
t.co/yE5uFPEof2

t.co/yE5uFPEof2
journals.plos.org/ploscompbiol...

journals.plos.org/ploscompbiol...
www.biorxiv.org/content/10.1...
www.biorxiv.org/content/10.1...


