
http://koo-lab.github.io

Excellent lineup of invited speakers across various scales of biology!
Deadline for abstract submission is coming up — Dec 2.
🔗 www.embl.org/about/info/c...
#EESAIBio @EMBLEvents

Excellent lineup of invited speakers across various scales of biology!
Deadline for abstract submission is coming up — Dec 2.
🔗 www.embl.org/about/info/c...
#EESAIBio @EMBLEvents

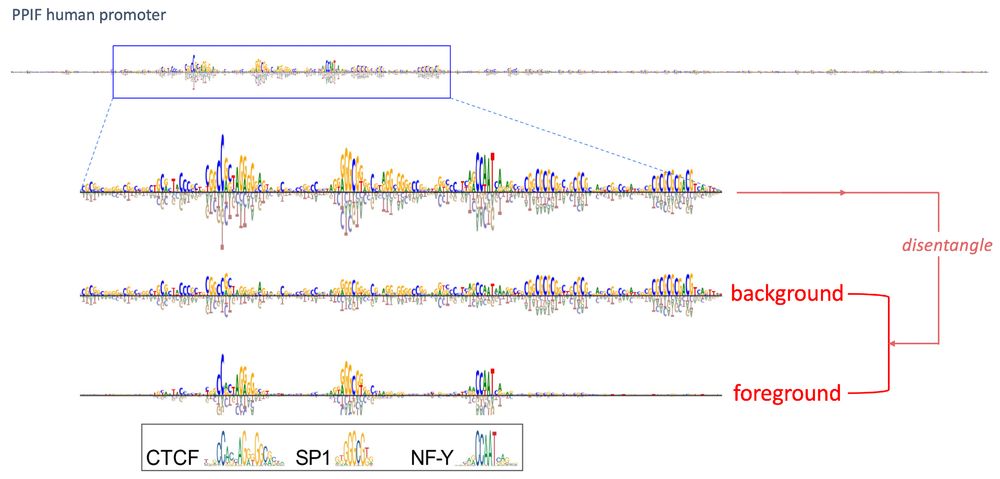
- single mutations -> no change
- double mutation -> CAAT box + new Inr
8/N

- single mutations -> no change
- double mutation -> CAAT box + new Inr
8/N

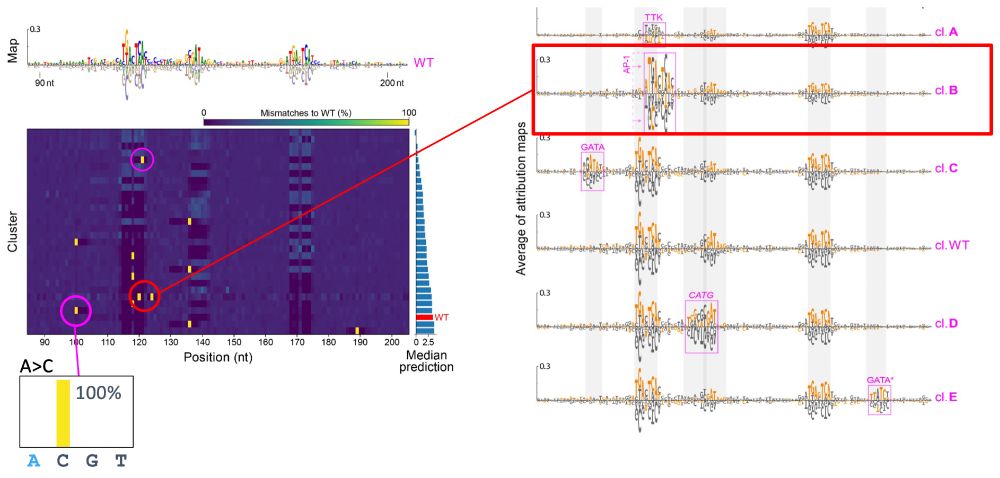



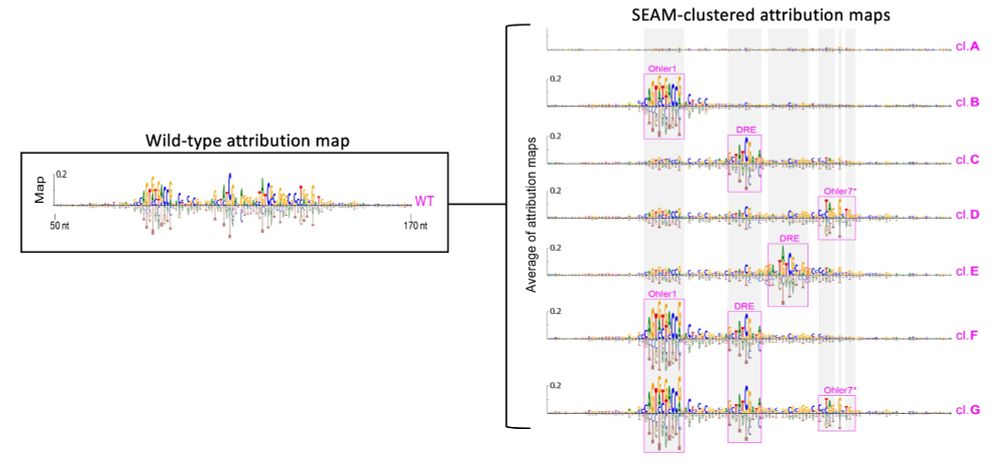
1) sample in a local region of sequence space via partial random mutagenesis
2) calculate attr maps to unveil the mechanisms
3) cluster attr maps based on shared mechanisms
4) cluster-based sequence analysis
2/N

1) sample in a local region of sequence space via partial random mutagenesis
2) calculate attr maps to unveil the mechanisms
3) cluster attr maps based on shared mechanisms
4) cluster-based sequence analysis
2/N
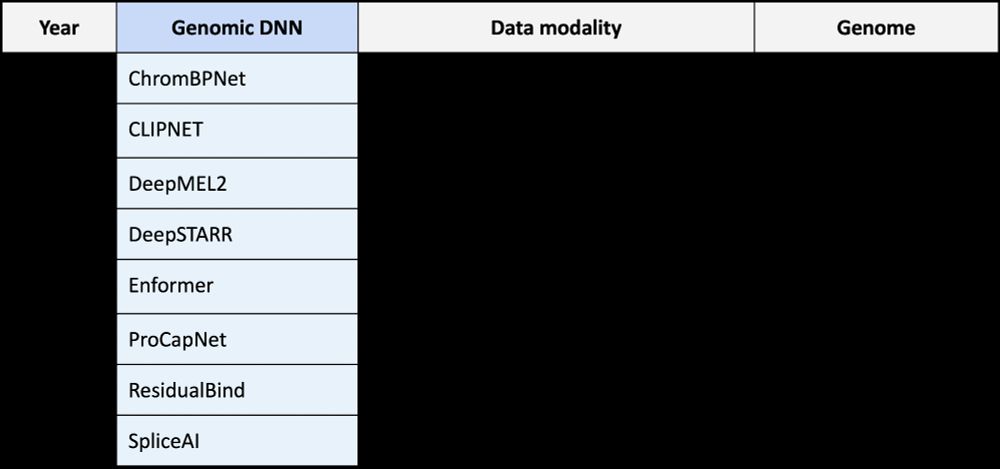
Excited to share SEAM: Systematic Explanation of Attribtuion-based Mechanisms. SEAM is an explainable AI method that dissects cis-regulatory mechanisms learned by seq2fun genomic deep learning models.
Led by @EESetiz
1/N 🧵👇

Excited to share SEAM: Systematic Explanation of Attribtuion-based Mechanisms. SEAM is an explainable AI method that dissects cis-regulatory mechanisms learned by seq2fun genomic deep learning models.
Led by @EESetiz
1/N 🧵👇
www.nobelprize.org/prizes/physi...
I have fond memories of my time in the Clarke lab, where I did my Honors Thesis on ultra low-field MRI w/ SQUIDs as an undergrad at UC Berkeley!

www.nobelprize.org/prizes/physi...
I have fond memories of my time in the Clarke lab, where I did my Honors Thesis on ultra low-field MRI w/ SQUIDs as an undergrad at UC Berkeley!







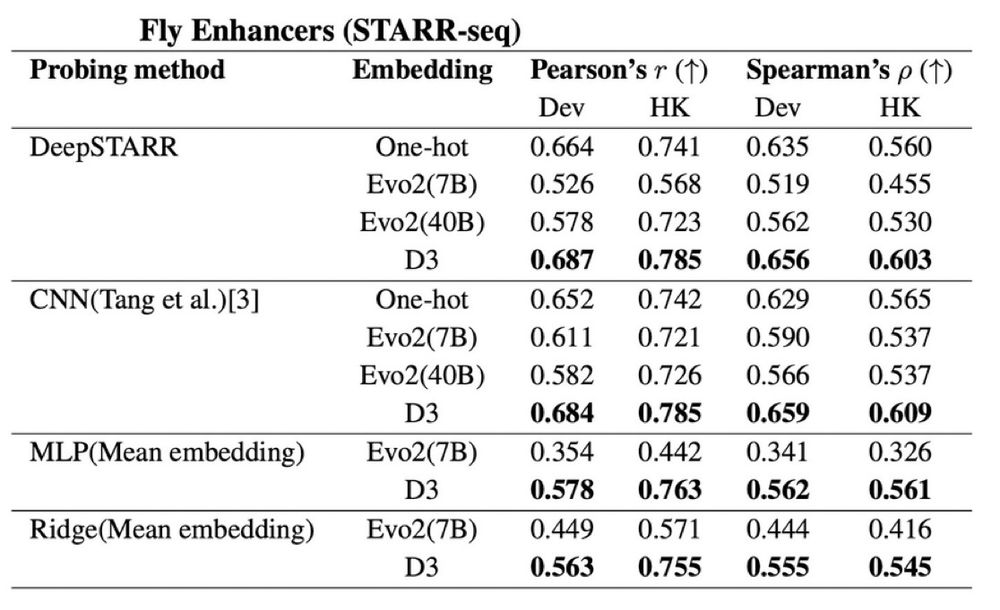
We compared probing strategies to assess how informative the pretrained representations are—benchmarking Evo2 vs D3 on Drosophila enhancer activity measured via STARR-seq.
Again, D3 outperforms Evo2 (and one-hot) across all probing methods!
We compared probing strategies to assess how informative the pretrained representations are—benchmarking Evo2 vs D3 on Drosophila enhancer activity measured via STARR-seq.
Again, D3 outperforms Evo2 (and one-hot) across all probing methods!

*Scaling* as the primary strategy with hopes of emergent properties is lazy.
Will the plan to fuse representations across mediocre (unimodal) foundation models work?!

*Scaling* as the primary strategy with hopes of emergent properties is lazy.
Will the plan to fuse representations across mediocre (unimodal) foundation models work?!
The result? D3-generated sequences are informative—they improve downstream supervised models, especially when paired with training tricks like EvoAug! (9/n)

The result? D3-generated sequences are informative—they improve downstream supervised models, especially when paired with training tricks like EvoAug! (9/n)

