
akdemirlab.github.io

EGFR FISH on post-Xenium slides showed >70% of these hubs localized to the nucleus and overlapped with EGFR copy number - unlike control gene FABP7 with similar expression but no hub formation.


EGFR FISH on post-Xenium slides showed >70% of these hubs localized to the nucleus and overlapped with EGFR copy number - unlike control gene FABP7 with similar expression but no hub formation.




We identified distinct cell states in IDH-wildtype vs mutant gliomas

We identified distinct cell states in IDH-wildtype vs mutant gliomas




For example, for p59, the short-read reconstruction suggested EGFR and CDK4 genes are on the same ecDNA circle.
However, Hi-C and ONT WGS-based reconstructions suggested these genes reside on two distinct circles.

For example, for p59, the short-read reconstruction suggested EGFR and CDK4 genes are on the same ecDNA circle.
However, Hi-C and ONT WGS-based reconstructions suggested these genes reside on two distinct circles.
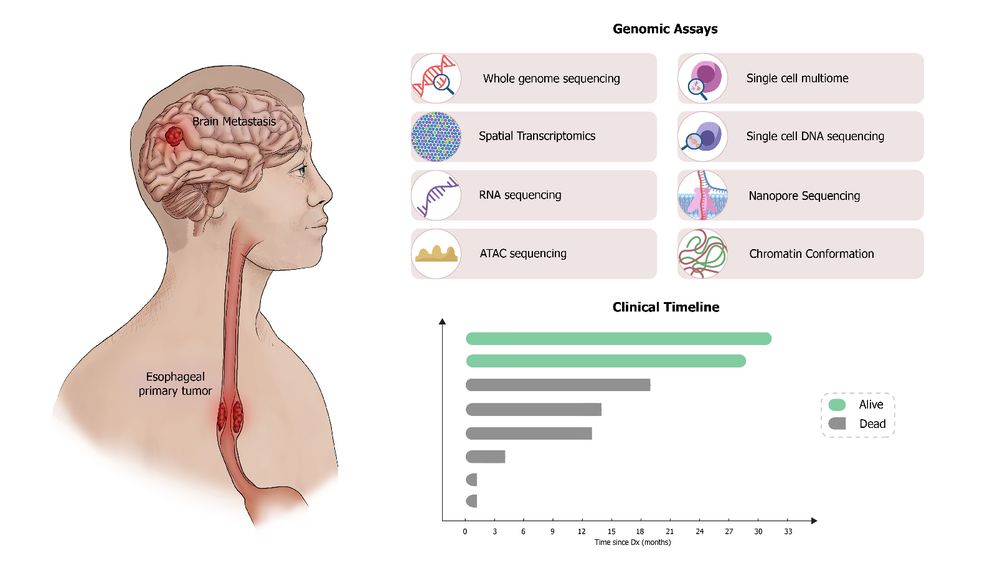
We used short- and long-read whole-genome sequencing plus Hi-C to reconstruct complex extrachromosomal DNA (ecDNA) structures. This revealed:
Tumors can harbor multiple ecDNA circles with distinct oncogenes.


We used short- and long-read whole-genome sequencing plus Hi-C to reconstruct complex extrachromosomal DNA (ecDNA) structures. This revealed:
Tumors can harbor multiple ecDNA circles with distinct oncogenes.
This was a great collaboration with Drs. Huse, Lang, @jesserdixon.bsky.social

This was a great collaboration with Drs. Huse, Lang, @jesserdixon.bsky.social









90% of EAC brain metastases harbor ERBB2 amplifications —MUCH higher than primary EAC (~20%).
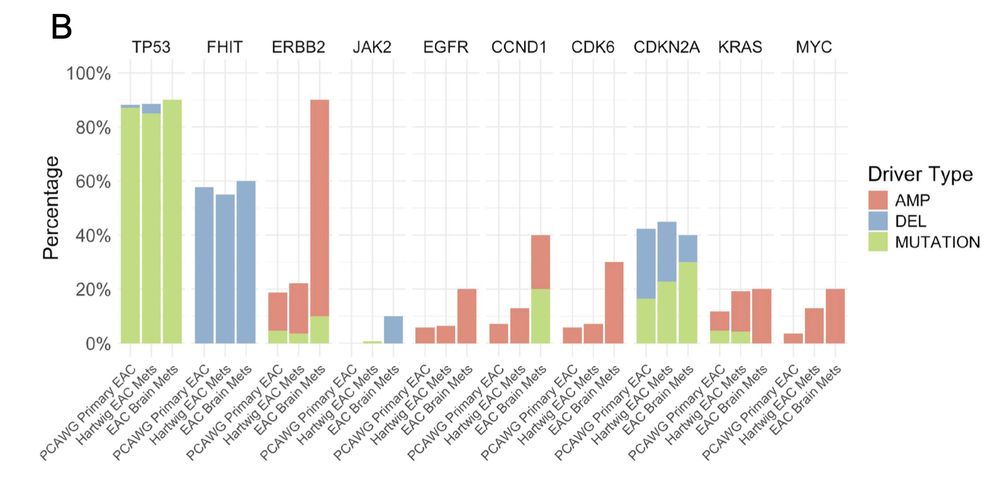
90% of EAC brain metastases harbor ERBB2 amplifications —MUCH higher than primary EAC (~20%).
It will be a long thread but I promise it's worth it!
#Oncology #BrainMets
It will be a long thread but I promise it's worth it!
#Oncology #BrainMets


