A Primer for Phylogenetic Causal Inference
with @oschwery.bsky.social & @pseudacris.bsky.social
Also, lots of new info on the website and more soon:
ssb2026.github.io
Follow for the latest updates!
@jembrown.bsky.social @systbiol.bsky.social
A Primer for Phylogenetic Causal Inference
with @oschwery.bsky.social & @pseudacris.bsky.social
Also, lots of new info on the website and more soon:
ssb2026.github.io
Follow for the latest updates!
@jembrown.bsky.social @systbiol.bsky.social


👀 Opportunities coming soon as I re-root my research program in plant ecophysiology and evolution at Wake 🌿 🌲🌱
#NewPI #plantecophys #botany @wfubiology.bsky.social
@wakeforest.bsky.social
👀 Opportunities coming soon as I re-root my research program in plant ecophysiology and evolution at Wake 🌿 🌲🌱
#NewPI #plantecophys #botany @wfubiology.bsky.social
@wakeforest.bsky.social
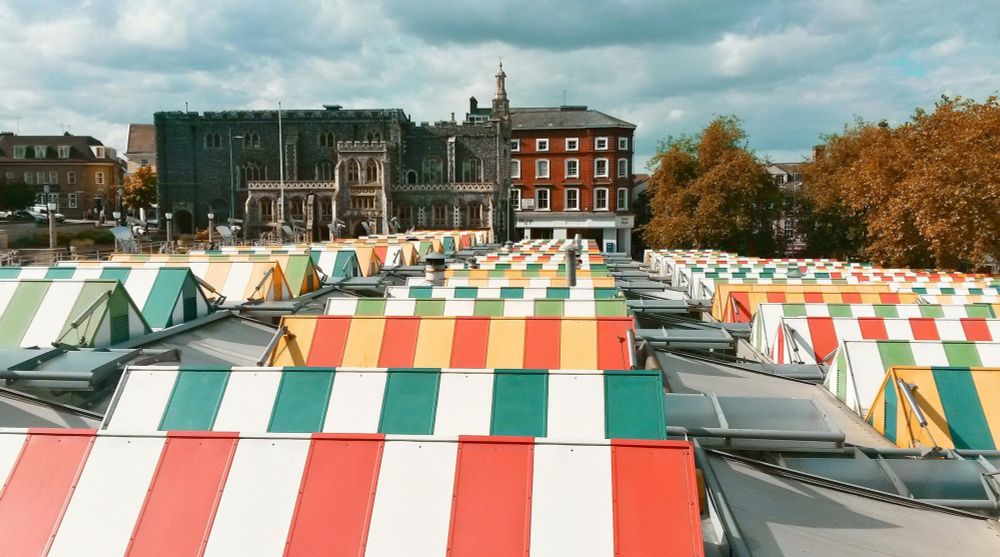


www.biorxiv.org/content/10.1...

www.biorxiv.org/content/10.1...

Very happy to share the published version of our work on natural Marchantia-Pseudomonas interactions.
Out now in Current Biology: www.cell.com/current-biol...
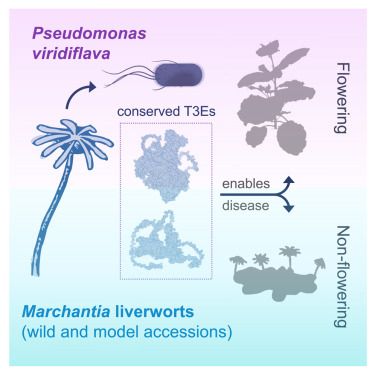
Very happy to share the published version of our work on natural Marchantia-Pseudomonas interactions.
Out now in Current Biology: www.cell.com/current-biol...


This research shatters the assumption that herbarium specimens are unsuitable for transcriptomics, successfully extracting mRNA from historical plant samples:
open.spotify.com/episode/1cqA...
This research shatters the assumption that herbarium specimens are unsuitable for transcriptomics, successfully extracting mRNA from historical plant samples:
open.spotify.com/episode/1cqA...







