We are building an AgentOps platform to observe, evaluate, and optimize enterprise AI agents and we are hiring. DM me if interested.

We are building an AgentOps platform to observe, evaluate, and optimize enterprise AI agents and we are hiring. DM me if interested.
NEMO again outperforms both the VAE-based and supervised approaches.

NEMO again outperforms both the VAE-based and supervised approaches.
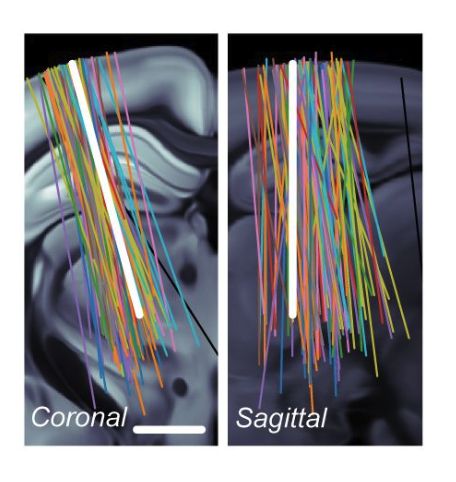
Without using any labels, NEMO's features align closely with anatomical regions and are consistent across labs.

Without using any labels, NEMO's features align closely with anatomical regions and are consistent across labs.
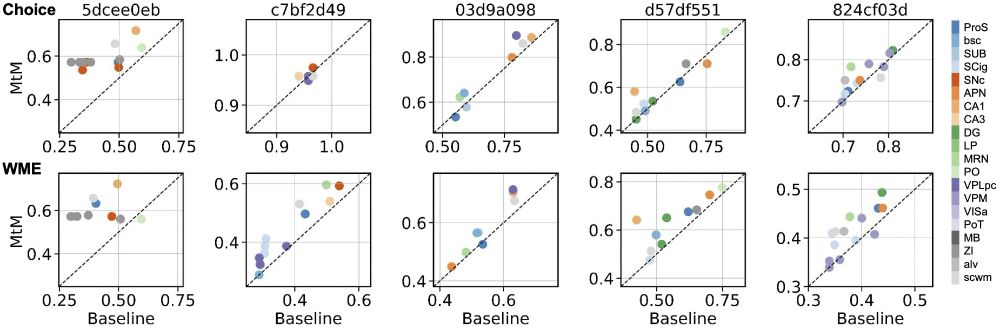
NEMO outperforms all baselines, including fully supervised models, with minimal fine-tuning.


NEMO outperforms all baselines, including fully supervised models, with minimal fine-tuning.
NEMO is trained to align ACGs and waveforms in a shared embedding space.

NEMO is trained to align ACGs and waveforms in a shared embedding space.
We use NEMO to characterize the electrophysiological diversity of cell-types across the entire mouse brain. 🐭 🧪 🧠
We use NEMO to characterize the electrophysiological diversity of cell-types across the entire mouse brain. 🐭 🧪 🧠
However, scaling laws are most useful when defined by performance as a function of model size, data, and compute. The POYO paper comes closest, showing scaling with both model and data size.
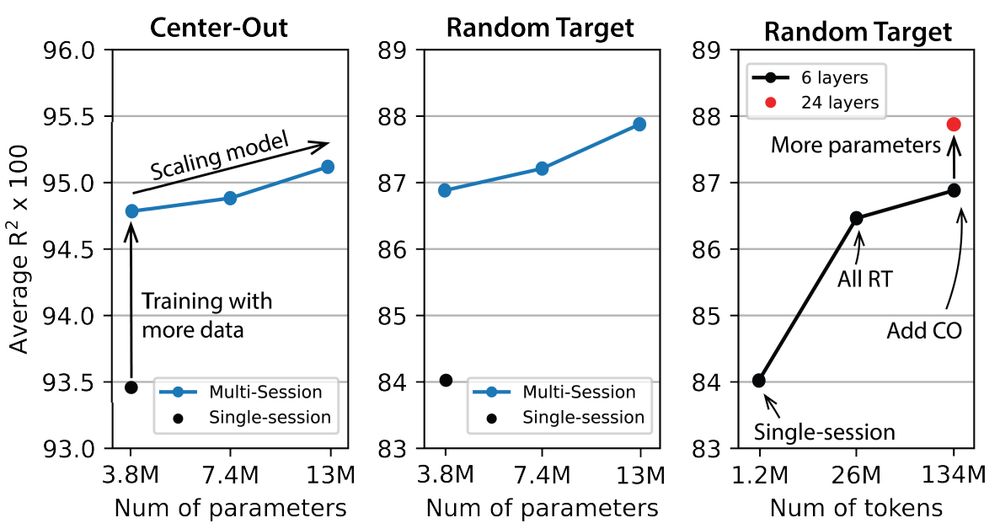
However, scaling laws are most useful when defined by performance as a function of model size, data, and compute. The POYO paper comes closest, showing scaling with both model and data size.




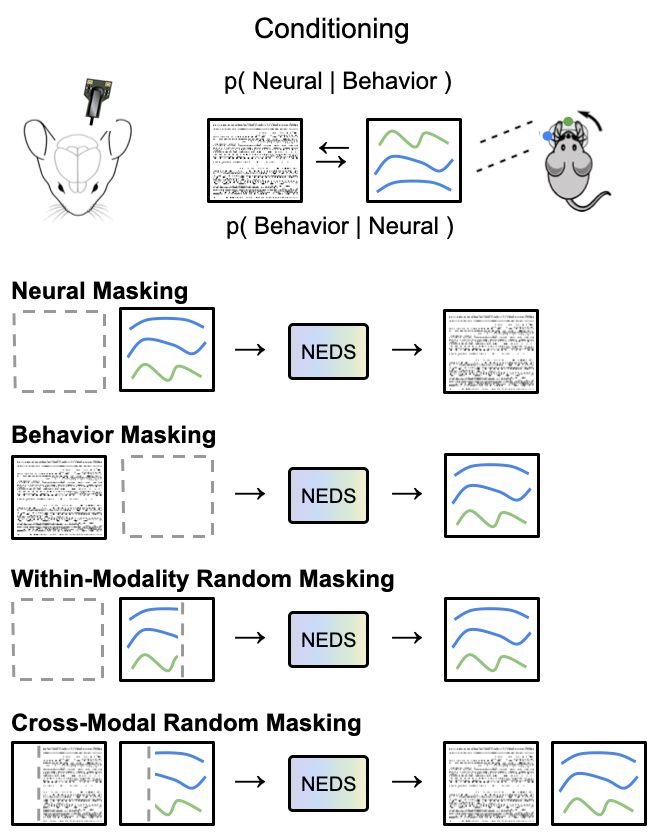
We train NEDS by masking out and reconstructing neural activity, behavior, within-modality, and across modalities.

We train NEDS by masking out and reconstructing neural activity, behavior, within-modality, and across modalities.
Trained on neural and behavioral data from 70+ mice, NEDS achieves state-of-the-art prediction of behavior (decoding) and neural responses (encoding) on held-out animals. 🐀
Trained on neural and behavioral data from 70+ mice, NEDS achieves state-of-the-art prediction of behavior (decoding) and neural responses (encoding) on held-out animals. 🐀




What will a foundation model for the brain look like? 🧠
We argue that it must be able to solve a diverse set of tasks across multiple brain regions and animals.
Check out our NeurIPS paper which introduces a multi-region, multi-animal, multi-task model arxiv.org/abs/2407.14668
What will a foundation model for the brain look like? 🧠
We argue that it must be able to solve a diverse set of tasks across multiple brain regions and animals.
Check out our NeurIPS paper which introduces a multi-region, multi-animal, multi-task model arxiv.org/abs/2407.14668

