Proteomics at Spanish National Centre for Cardiovascular Research
🔬 Captures multiple PTM layers (trypsin efficiency, zonal changes, PTMs, hypermodifications)
⚡ Advanced artifact control
🐭 Uncovering novel oxidative modifications
🔬 Captures multiple PTM layers (trypsin efficiency, zonal changes, PTMs, hypermodifications)
⚡ Advanced artifact control
🐭 Uncovering novel oxidative modifications
We're looking forward to exploring its full potential in the coming weeks.
📸👇
#Proteomics #CNICproteomics #Cardiovasculardeseases #Opentrons #LabAutomation

We're looking forward to exploring its full potential in the coming weeks.
📸👇
#Proteomics #CNICproteomics #Cardiovasculardeseases #Opentrons #LabAutomation
#Proteomics #PTMs #SEBBM2025 #MassSpectrometry #CNICproteomics #Cardiovasculardeseases
#Proteomics #PTMs #SEBBM2025 #MassSpectrometry #CNICproteomics #Cardiovasculardeseases
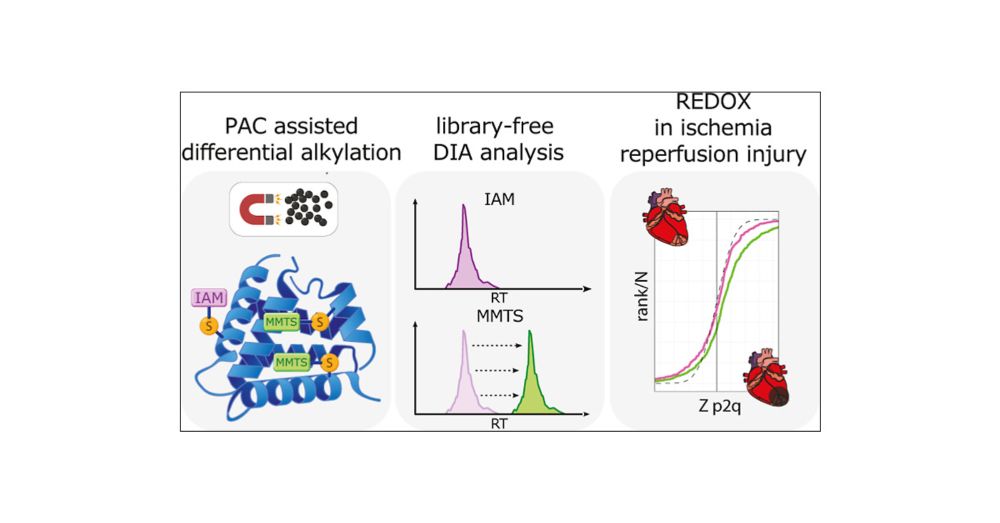
👉 Published in Analytical Chemistry: pubs.acs.org/doi/full/10....
Contact us if you want to get the protocol 🗒️
👉 Published in Analytical Chemistry: pubs.acs.org/doi/full/10....
Contact us if you want to get the protocol 🗒️
Stay tuned for updates, publications, and opportunities to engage with our team
#Proteomics #MassSpectrometry #OpenSearch #PTMs #CNICproteomics
Stay tuned for updates, publications, and opportunities to engage with our team
#Proteomics #MassSpectrometry #OpenSearch #PTMs #CNICproteomics
1️⃣ Map the landscape of PTMs in complex biological systems.
2️⃣ Provide insights into the molecular mechanisms underlying cardiovascular diseases.
1️⃣ Map the landscape of PTMs in complex biological systems.
2️⃣ Provide insights into the molecular mechanisms underlying cardiovascular diseases.


