
Kudos to co-coordinators @miriamlisci.bsky.social and @juliensluneau.bsky.social, and to the fantastic team of 19 volunteers at the bars 🥳!
#PINT25 #scicomm




Kudos to co-coordinators @miriamlisci.bsky.social and @juliensluneau.bsky.social, and to the fantastic team of 19 volunteers at the bars 🥳!
#PINT25 #scicomm
(7/8)

(7/8)
Interestingly, this domain is only present in insects and most vertebrates.
(5/8)

Interestingly, this domain is only present in insects and most vertebrates.
(5/8)
#SIRT7 binding is optimal for targeting #H3K36ac, and it can only access #H3K18ac through partial disengagement from the nucleosome and DNA bending.
(4/8)

- A unique N-terminal nucleosome-binding domain, which places the #SIRT7 catalytic domain by the H3 tail exit site
- Extensive contacts with the two DNA gyres!
(3/8)

- A unique N-terminal nucleosome-binding domain, which places the #SIRT7 catalytic domain by the H3 tail exit site
- Extensive contacts with the two DNA gyres!
(3/8)
This allowed us to stabilize specific enzyme:substrate complexes efficiently for #cryoEM.
(2/8)

This allowed us to stabilize specific enzyme:substrate complexes efficiently for #cryoEM.
(2/8)
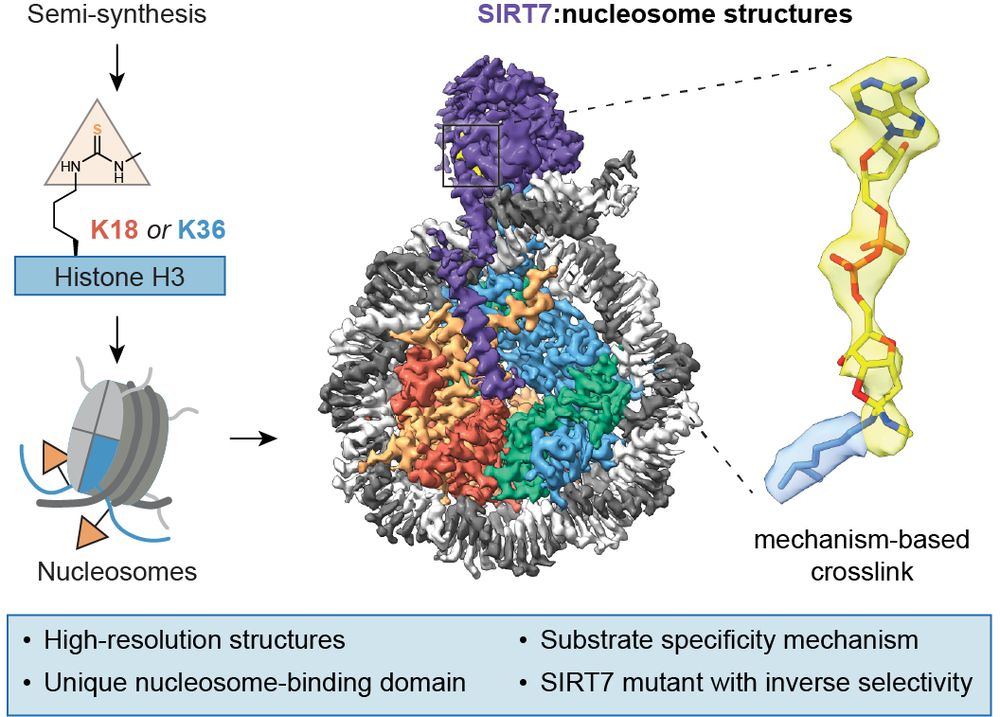
We just published mechanism-based #cryoEM structures of #SIRT7 on nucleosomes to understand its activity 👇
www.nature.com/articles/s41...
(1/8) #ChemBio #ChemSky
We just published mechanism-based #cryoEM structures of #SIRT7 on nucleosomes to understand its activity 👇
www.nature.com/articles/s41...
(1/8) #ChemBio #ChemSky


