The residues aren't directly adjacent, suggesting that the model isn't simply memorizing training data:

The residues aren't directly adjacent, suggesting that the model isn't simply memorizing training data:


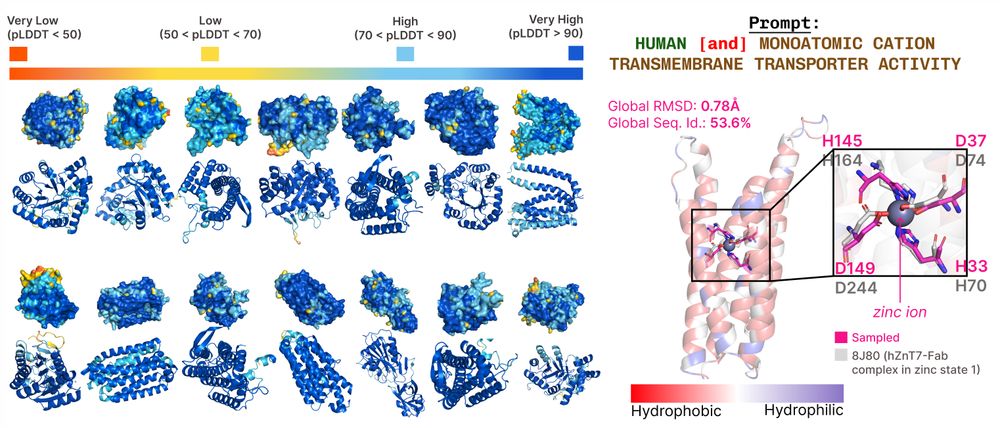
For inference, we can sample latent embeddings & use frozen sequence/structure decoders to get all-atom structure:

For inference, we can sample latent embeddings & use frozen sequence/structure decoders to get all-atom structure:
bit.ly/plaid-proteins
🧵
bit.ly/plaid-proteins
🧵

